|
Middle East respiratory syndrome coronavirus (MERS-CoV)
originated in bats and spread to humans via zoonotic
transmission from camels.
Researchers analyzed the evolution of the spike (S)
gene in betacoronaviruses (betaCoVs) isolated from
different mammals, in bat coronavirus populations,
as well as in MERS-CoV strains from the current
outbreak.
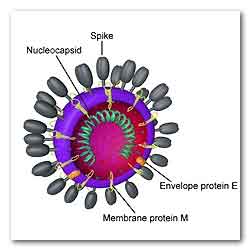
"Coronavirus virion" di Belouzard, et al -
www.ncbi.nlm.nih.gov/pmc/articles/PMC3397359/.
Results indicated several positively selected sites
located in the region comprising the two heptad
repeats (HR1 and HR2) and their linker.
Two sites (R652 and V1060) were positively selected
in the betaCoVs phylogeny and correspond to
mutations associated with expanded host range in
other coronaviruses.
During the most recent evolution of MERS-CoV,
adaptive mutations in the HR1 (Q/R/H1020) arose in
camels or in a previous host and spread to humans.
Researchers determined that different residues at
position 1020 establish distinct inter- and
intra-helical interactions and affect the stability
of the six-helix bundle formed by the HRs.
A similar effect on stability was observed for a
nearby mutation (T1015N) that increases MERS-CoV
infection efficiency in vitro. Data herein indicate
that the heptad repeat region was a major target of
adaptive evolution in MERS-CoV-related viruses;
these results are relevant for the design of fusion
inhibitor peptides with antiviral function.
For more information
The heptad repeat region is a major selection target
in MERS-CoV and related coronaviruses
Diego Forni, Giulia Filippi, Rachele Cagliani, Luca
De Gioia, Uberto Pozzoli, Nasser Al-Daghri, Mario
Clerici e Manuela Sironi
http://www.nature.com/articles/srep14480
MDN |