|
The finding of key genetic switches called
super-enhancers, involved in regulating the human
immune system opens the door to new research into
autoimmune disorders such as inflammatory bowel
disease or rheumatoid arthritis.
The immune system has a complex, delicately
orchestrated balance. White blood cells called CD4 T
cells can mature to become many types of T cells,
each of which has a distinct function. Some activate
immune responses; others constrain immune responses.
When the system is out of balance, uncontrolled
reactions can lead to attacks against the body’s own
cells and tissues and cause autoimmune disease. Many
different tissues can be affected. For example,
joints become swollen and inflamed in rheumatoid
arthritis, and the brain and spinal cord are damaged
in multiple sclerosis.
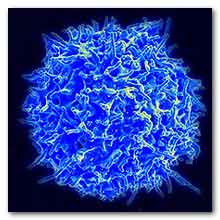
Scanning electron micrograph of a human T cell from the immune system of a healthy donor. Source: National Institute of Allergy and Infectious Diseases (NIAID).
Autoimmune diseases often run in families. However,
identifying susceptibility genes has been a
challenge. In most cases, a mix of genetic and
environmental factors is at play. The genetic
variants tied to these diseases tend to lie in
regions involved in regulating genes, rather than in
genes themselves.
A research team led by Drs. Golnaz Vahedi and John
J. O’Shea at NIH’s National Institute of Arthritis
and Musculoskeletal and Skin Diseases (NIAMS)
investigated the role of a recently discovered type
of genetic regulatory element called super-enhancers,
or stretch-enhancers (SE). Earlier work in other
laboratories—including that of NIH Director Dr.
Francis S. Collins—showed that SEs are especially
powerful switches that control genes important for
cell identity. A large number of disease-associated
genetic alterations have been found to fall within
SEs.
The team searched the genome of T cells for regions
bound by a protein called histone acetyltransferase
p300, which marks DNA segments that carry SEs. As
reported online in Nature on February 16, 2015, the
scientists found several hundred genes associated
with SEs.
The dominant gene class associated with SEs in T
cells encoded cytokines and cytokine receptors.
These allow T cells to communicate with other cells
and coordinate the immune response.
A large fraction of variants previously associated
with rheumatoid arthritis and other autoimmune
diseases also localized to T cell SEs. However, the
greatest SE enrichment in T cells was associated
with the gene for BACH2, which has been previously
associated with rheumatoid arthritis and other
autoimmune diseases.
Recently, NIH scientists found that a major function
of BACH2 is controlling the activation of T cells.
When the scientists exposed human T cells to
tofacitinib—a drug used to treat rheumatoid
arthritis—the activities of many genes controlled by
SEs were preferentially affected. However, BACH2
levels were unchanged.
This result suggests that tofacitinib may act on SEs
independent of BACH2 to alter the activities of
important T cell genes.
“Three types of data—the genetics of rheumatoid
arthritis, a genomic feature of T cells, and the
pharmacological effects of a rheumatoid arthritis
drug—are all pointing to the importance of
super-enhancers,” Vahedi says. “These regions are
where we plan to search for insights into the
mechanisms that underlie rheumatoid arthritis and
other autoimmune diseases, and for novel therapeutic
targets for these conditions.”
For more information
Super-enhancers delineate disease-associated
regulatory nodes in T cells.
Vahedi G, Kanno Y, Furumoto Y, Jiang K, Parker SC,
Erdos MR, Davis SR, Roychoudhuri R, Restifo NP,
Gadina M, Tang Z, Ruan Y, Collins FS, Sartorelli V,
O'Shea JJ.
Nature. 2015 Feb 16. doi: 10.1038/nature14154. [Epub
ahead of print]. PMID: 25686607.
MDN |